Replacing the Magrittr Pipe With the Native R Pipe
By Katie Press
June 19, 2022
What’s the deal with the new pipe?
R version 4.0+ has a “native” pipe |> that does not rely on the magrittr dependency. Apparently it is supposed to be faster than the magrittr pipe, which does a lot more “behind the scenes”. You can read more about it in
R for Data Science. There are also a couple of other blog posts out there on the subject. I decided to give it a go since I like to try new things, so I decided to run a bunch of my old code, just replacing the old pipe with the native pipe to see what would happen. Surprisingly I had very few issues, but I thought I would write this blog posts describing the couple times where it did fail, and how I fixed it.
knitr::opts_chunk$set(echo = TRUE)
library(esvis)
library(tidyverse)
library(rvest)
library(janitor)
First of all, you can change your shortcut to the native pipe operator through the global settings under “code” like so:
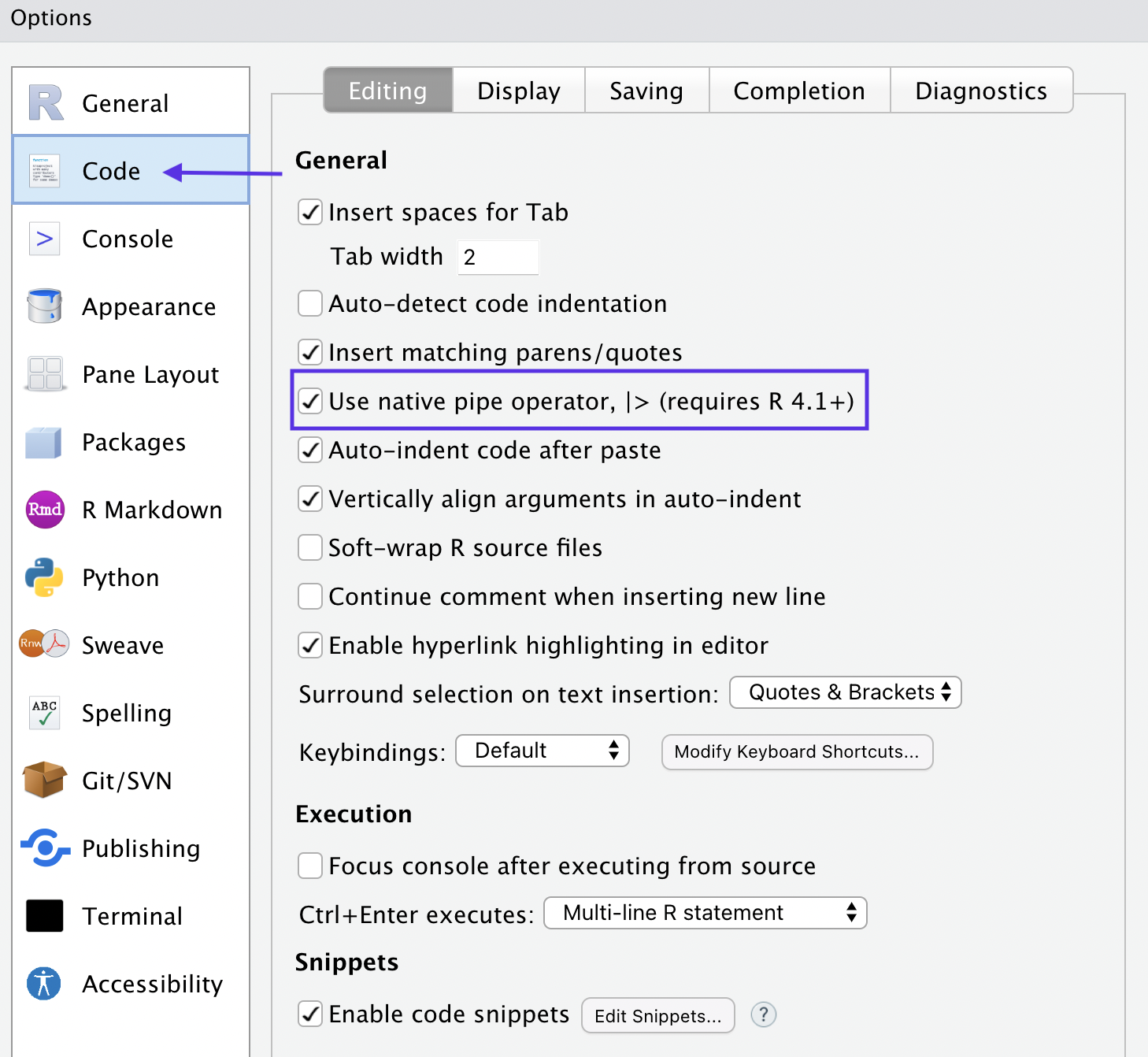
There are some differences to be aware of when using the native pipe - primarily that it doesn’t work with dot notation. You have to use an anonymous function instead, which I will illustrate in the three examples in this post.
Example 1: mapping to clean names
I don’t want to load all the data necessary for this example, so here’s a quick list of dataframes to use.
data_list <- list(
data1 = tibble(Var1 = c("a", "b", "c"), `Variable #2` = c(1,2,3)),
data2 = tibble(Var1 = c("a", "b", "c"), `Variable #2` = c(1,2,3))
)
This code did not work when I replaced it with the native pipe.
data_list %>%
map( ~.x %>% clean_names)
Again, it just needed to be replaced with an anonymous function. This time I’ve used x instead of the dot so you can see it still works.
data_list |>
map(\(x) clean_names(x))
## $data1
## # A tibble: 3 × 2
## var1 variable_number_2
## <chr> <dbl>
## 1 a 1
## 2 b 2
## 3 c 3
##
## $data2
## # A tibble: 3 × 2
## var1 variable_number_2
## <chr> <dbl>
## 1 a 1
## 2 b 2
## 3 c 3
Example 2: The native pipe doesn’t work with split and dot notation
The first code failure is from a file I created for someone who asked how they could map over several variables when testing effect sizes. This is really just an example to show the features of purrr. I’m using example data from the esvis package.
data(star)
star <- star |>
select(condition, sex, reading) |>
gather(variable, value, - reading)
This was my original code, and when I replaced the magrittr pipe with the native pipe it didn’t work.
star %>%
split(.$variable) %>%
map_df(. %>% hedg_g(., reading ~ value), .id = "variable")
The native pipe does not allow the use of the period placeholder with the dollar sign, or with brackets for that matter which we’ll see in a minute. But the tilde works to split by variable, and then I just had to replace the dot notation with an anonymous function instead. It doesn’t have to use the dot, you could replace it with x or whatever name you want.
star |>
split(~variable) |>
map_df(\(.) hedg_g(., reading ~ value), .id = "variable")
## # A tibble: 8 × 5
## variable value_ref value_foc coh_d hedg_g
## <chr> <chr> <chr> <dbl> <dbl>
## 1 condition reg reg_aid 0.0258 0.0258
## 2 condition reg small -0.159 -0.159
## 3 condition reg_aid reg -0.0258 -0.0258
## 4 condition reg_aid small -0.186 -0.186
## 5 condition small reg 0.159 0.159
## 6 condition small reg_aid 0.186 0.186
## 7 sex boy girl -0.193 -0.193
## 8 sex girl boy 0.193 0.193
Example 3: Using the native pipe while accessing data with brackets
Remember I said the base pipe would not allow the dot notation with brackets? This is an example of the code I tried to run which didn’t work when I just replaced the pipe.
wiki <- "https://en.wikipedia.org/wiki/Dow_Jones_Industrial_Average"
djia <- wiki %>%
read_html() %>%
html_nodes("table") %>%
.[[2]] %>%
html_table(fill = TRUE) %>%
clean_names()
This one took me a minute to figure out. I had to encase the entire line in parentheses, use the anonymous function, and then add extra empty parentheses at the end of the line. But it worked!
wiki |>
read_html() |>
html_nodes("table") |>
(\(.) .[[2]])() |>
html_table(fill = TRUE) |>
clean_names()
## # A tibble: 30 × 7
## company exchange symbol industry date_added notes index_weighting
## <chr> <chr> <chr> <chr> <chr> <chr> <chr>
## 1 3M NYSE MMM Conglomera… 1976-08-09 "As … 2.88%
## 2 American Express NYSE AXP Financial … 1982-08-30 "" 3.56%
## 3 Amgen NASDAQ AMGN Biopharmac… 2020-08-31 "" 4.88%
## 4 Apple NASDAQ AAPL Informatio… 2015-03-19 "" 3.15%
## 5 Boeing NYSE BA Aerospace … 1987-03-12 "" 3.40%
## 6 Caterpillar NYSE CAT Constructi… 1991-05-06 "" 4.19%
## 7 Chevron NYSE CVX Petroleum … 2008-02-19 "Als… 3.05%
## 8 Cisco NASDAQ CSCO Informatio… 2009-06-08 "" 1.00%
## 9 Coca-Cola NYSE KO Drink indu… 1987-03-12 "Als… 1.28%
## 10 Disney NYSE DIS Broadcasti… 1991-05-06 "" 2.32%
## # … with 20 more rows
Fira code and ligature fonts
Now I’ll write some code just to show you how cool it looks if you use the Fira Code font and enable ligatures. My blog Rmarkdown file will not show the ligatures by default, so the code chunk below is how it would look without Fira Code.
djia |>
filter(exchange == "NYSE" & company != "3M") |>
filter(date_added >= "2009-01-01") -> new_djia
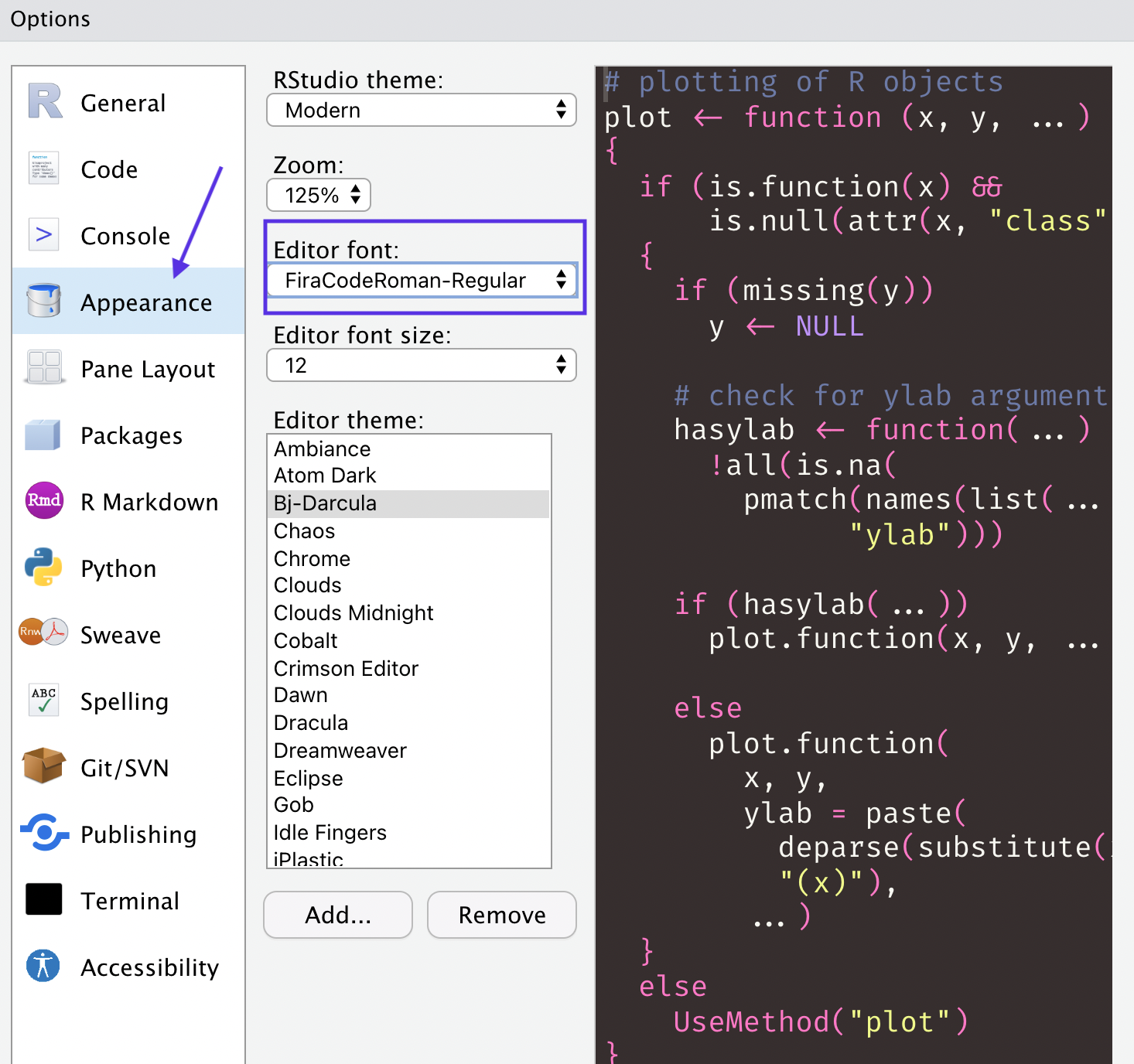
And here’s what it looks like in my RStudio console. Pretty cool!

You can do this yourself if you download Fira Code which you can get from Google Fonts. Then go to the global options and change the font settings. Note that you will most likely have to restart RStudio before the font will show up in your list, so don’t worry if you don’t see it there right away.
- Posted on:
- June 19, 2022
- Length:
- 5 minute read, 996 words
- See Also: